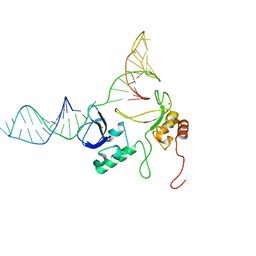1. F1F2
2. DNA (45-MER)
Sample #1
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 1 | PARP-1 1-214 | [U-15N; U-13C; U-2H] | | 0.2 mM |
|---|
| 2 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 3 | TRIS | [U-2H] | | 50 mM |
|---|
| 4 | DTT | [U-2H] | | 1 mM |
|---|
| 5 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 6 | H2O | natural abundance | solvent | 95 % |
|---|
| 7 | D2O | [U-2H] | solvent | 5 % |
|---|
Sample #2
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H (approx. 70%),15N,13C]. This sample was only used to confirm
assignments by checking patterns of peaks; no numerical shift data from
experiments with this sample contributed to the deposition (since isotope
shifts could differ).
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 8 | PARP-1 1-214 | [U-15N; U-13C; U-70% 2H] | | 0.2 mM |
|---|
| 9 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 10 | TRIS | [U-2H] | | 50 mM |
|---|
| 11 | DTT | [U-2H] | | 1 mM |
|---|
| 12 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 13 | H2O | natural abundance | solvent | 95 % |
|---|
| 14 | D2O | [U-2H] | solvent | 5 % |
|---|
Sample #3
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 15 | PARP-1 1-214 | [U-98% 2H; U-98% 15N] | | 0.2 mM |
|---|
| 16 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 17 | TRIS | [U-2H] | | 50 mM |
|---|
| 18 | DTT | [U-2H] | | 1 mM |
|---|
| 19 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 20 | H2O | natural abundance | solvent | 95 % |
|---|
| 21 | D2O | [U-2H] | solvent | 5 % |
|---|
Sample #4
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C], back-labeled with [1H,13C] in the delta-methyl groups of
Ile and all methyl groups of Leu and Val residues, using sodium [4-13C, 3,3-2H2]
alpha-ketobutyrate and sodium [3- 2H, 4,4'-13C2] alpha-ketoisovalerate as
precursors to maximize protonation of methyl groups, for use in NOE experiments;
sodium [3-2H, 4,4'-13C2] alpha-ketoisovalerate was prepared from sodium
[4,4'-13C2] alpha-ketoisovalerate by exchange with 2H2O at pH 12.5 and 45 C for
3 hrs.
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 22 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 23 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 24 | TRIS | [U-2H] | | 50 mM |
|---|
| 25 | DTT | [U-2H] | | 1 mM |
|---|
| 26 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 27 | H2O | natural abundance | solvent | 95 % |
|---|
| 28 | D2O | [U-2H] | solvent | 5 % |
|---|
Sample #5
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C], back-labeled with [1H,13C] in the delta-methyl groups of
Ile and all methyl groups of Leu and Val residues, using sodium [3,3-2H2,13C4]
alpha-ketobutyrate and sodium [3- 2H,13C5] alpha-ketoisovalerate as precursors
to produce linear chains of 13C in the sidechains of Val and Leu, for use in
assignment experiments to link methyl signals to C-alpha signals.
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 29 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 30 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 31 | TRIS | [U-2H] | | 50 mM |
|---|
| 32 | DTT | [U-2H] | | 1 mM |
|---|
| 33 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 34 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #6
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C], back-labeled with [1H,13C] in the methyl groups of Met
residues in addition to Ile, Leu and Val methyl groups as in sample 4a.
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 35 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 36 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 37 | TRIS | [U-2H] | | 50 mM |
|---|
| 38 | DTT | [U-2H] | | 1 mM |
|---|
| 39 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 40 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #7
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C]; back-labeled with [1H,13C] in the methyl groups of Ile,
Leu and Val methyl groups as in sample 4a and [1H,13C,15N] Phe residues.
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 41 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 42 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 43 | TRIS | [U-2H] | | 50 mM |
|---|
| 44 | DTT | [U-2H] | | 1 mM |
|---|
| 45 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 46 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #8
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Sortase ligated, block-labelled sample. (N.B. residues 103 and 104 of WT
sequence deleted, additional residues LPETGGG inserted between residues 102 and
105; this sample was not used for making any assignments of residues in this
region, which is in the flexible linker between domains).
Labelling for residues 1-102 (and LPET of insertion):
uniform [1H,12C,15N]
Labelling for residues 105-214 (and GGG of insertion):
[2H,15N,13C] back-labeled with [1H,13C] Ile, Leu Val and Met methyl groups
labeled as in sample 5
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 47 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 48 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 49 | TRIS | [U-2H] | | 50 mM |
|---|
| 50 | DTT | [U-2H] | | 1 mM |
|---|
| 51 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 52 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #9
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Sortase ligated, block-labelled sample. (N.B. residues 103 and 104 of WT
sequence deleted, additional residues LPETGGG inserted between residues 102 and
105; this sample was not used for making any assignments of residues in this
region, which is in the flexible linker between domains).
Labelling for residues 1-102 (and LPET of insertion):
uniform [1H,12C,15N]
Labelling for residues 105-214 (and GGG of insertion):
[2H,15N,13C] back labeled with [1H,13C] Met methyl groups as in sample 5 and
[13C,15N,1H] Arg residues
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 53 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 54 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 55 | TRIS | [U-2H] | | 50 mM |
|---|
| 56 | DTT | [U-2H] | | 1 mM |
|---|
| 57 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 58 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #10
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Sortase ligated, block-labelled sample. (N.B. residues 103 and 104 of WT
sequence deleted, additional residues LPETGGG inserted between residues 102 and
105; this sample was not used for making any assignments of residues in this
region, which is in the flexible linker between domains).
Labelling for residues 1-102 (and LPET of insertion):
[2H,15N,13C] back-labeled with Ile, Leu and Val methyl groups labeled as in
sample 4a and [1H,13C,15N] Arg residues
Labelling for residues 105-214 (and GGG of insertion):
uniform [1H,12C,15N]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 59 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 60 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 61 | TRIS | [U-2H] | | 50 mM |
|---|
| 62 | DTT | [U-2H] | | 1 mM |
|---|
| 63 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 64 | H2O | natural abundance | solvent | 95 % |
|---|
| 65 | D2O | [U-2H] | solvent | 5 % |
|---|
Sample #11
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Sortase ligated, block-labelled sample. (N.B. residues 103 and 104 of WT
sequence deleted, additional residues LPETGGG inserted between residues 102 and
105; this sample was not used for making any assignments of residues in this
region, which is in the flexible linker between domains).
Labelling for residues 1-102 (and LPET of insertion):
[2H,15N,13C] back-labeled with Ile, Leu and Val methyl groups labeled as in
sample 4a and [1H,13C,15N] Phe residues
Labelling for residues 105-214 (and GGG of insertion):
uniform [1H,12C,15N]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 66 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 67 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 68 | TRIS | [U-2H] | | 50 mM |
|---|
| 69 | DTT | [U-2H] | | 1 mM |
|---|
| 70 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 71 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #12
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Sortase ligated, block-labelled sample. (N.B. residues 103 and 104 of WT
sequence deleted, additional residues LPETGGG inserted between residues 102 and
105; this sample was not used for making any assignments of residues in this
region, which is in the flexible linker between domains).
Labelling for residues 1-102 (and LPET of insertion):
[2H,15N,13C] back-labeled with Ile, Leu and Val methyl groups labeled as in
sample 4a
Labelling for residues 105-214 (and GGG of insertion):
uniform [2H,12C,15N] back-labeled with [1H,15N,13C] Pro
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 72 | PARP-1 1-214 | see Sample details section | | 0.2 mM |
|---|
| 73 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 74 | TRIS | [U-2H] | | 50 mM |
|---|
| 75 | DTT | [U-2H] | | 1 mM |
|---|
| 76 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 77 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #13
Solvent system 100% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 78 | PARP-1 1-214 | [U-15N; U-13C; U-2H] | | 0.2 mM |
|---|
| 79 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 80 | TRIS | [U-2H] | | 50 mM |
|---|
| 81 | DTT | [U-2H] | | 1 mM |
|---|
| 82 | ZnSO4 | natural abundance | | 0.1 mM |
|---|
| 83 | D2O | [U-2H] | solvent | 100 % |
|---|
Sample #14
Solvent system 95% H2O/5% D2O, Pressure 1 atm, Temperature 273 K, pH 7.2, Details Uniform [2H,15N,13C]
| # | Name | Isotope labeling | Type | Concentration |
|---|
| 84 | PARP-1 1-214 | [U-15N; U-13C; U-2H] | | 0.2 mM |
|---|
| 85 | DNA (45-MER) | natural abundance | | 0.2 mM |
|---|
| 86 | TRIS | [U-2H] | | 50 mM |
|---|
| 87 | DTT | [U-2H] | | 1 mM |
|---|
| 88 | ZnSO4 | natural abundance | | 0.1 uM |
|---|
| 89 | sodium chloride | natural abundance | | 200 mM |
|---|
| 90 | H2O | natural abundance | solvent | 95 % |
|---|
| 91 | D2O | [U-2H] | solvent | 5 % |
|---|